|
To date, fluoride-resistant strains have been created for several streptococci, including S. To better understand the mechanism of microbial fluoride resistance, researchers also created fluoride-resistant strains in the laboratory by selecting colonies that were able to grow in the presence of 400–600 ppm fluoride. Fluoride-resistant strains have been discovered in several clinical studies, where fluoride-resistant Streptococcus mutans colonies were recovered from xerostomia patients who had been treated with gels containing a high concentration of NaF. One of the consequences of the widespread and prolonged application of fluoride is the risk of the development of fluoride resistance in microorganisms. It is known that fluoride prevents caries through a dual-mode of action it not only inhibits the demineralization and enhances the remineralization of dental hard tissues but also affects bacterial growth and microbial metabolic activity by the inhibition of enolase and ATPase. While the application of fluoride containing products has markedly reduced caries, the exact mechanism of fluoride in caries prevention is still not completely understood. Since its discovery as a preventive agent in 1931, fluoride has been widely used in many caries-preventive products and in many countries. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.Ĭompeting interests: The authors have declared that no competing interests exist.įluoride is still the most effective caries-preventive agent. 81371135) and "Twelfth Five-Year" National Science and Technology Support Project (20911BAZ03171). This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are creditedĭata Availability: All relevant data are within the paper and its supporting information files.įunding: This research was supported by the National Natural Science Foundation of China (No. Received: DecemAccepted: FebruPublished: April 9, 2015Ĭopyright: © 2015 Liao et al. PLoS ONE 10(4):Īcademic Editor: Justin Merritt, Oregon Health and Science University, UNITED STATES (2015) Identification and Functional Analysis of Genome Mutations in a Fluoride-Resistant Streptococcus mutans Strain. These findings can provide new insights into the mechanism of microbial fluoride resistance.Ĭitation: Liao Y, Chen J, Brandt BW, Zhu Y, Li J, van Loveren C, et al. In summary, using WGS sequencing, we were able to uncover genetic changes in the genome of a fluoride-resistant strain. 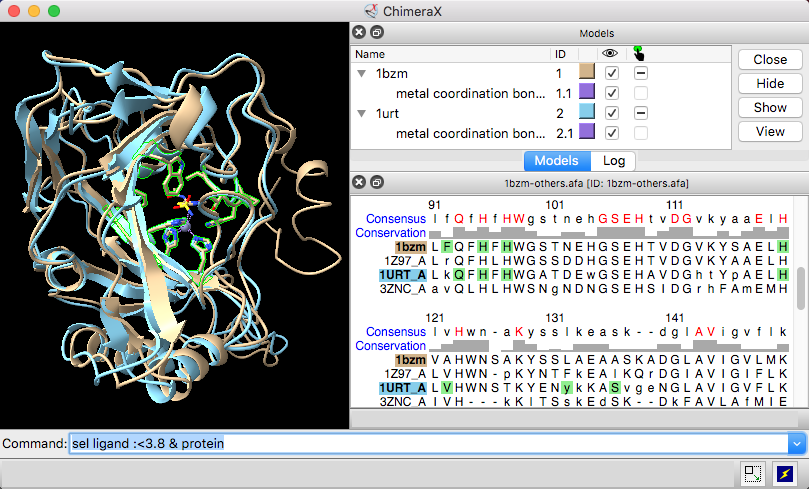
No difference in expression was found for the other SNP-containing genes. The promoter region of this gene contained a SNP. In addition, one gene, which codes for a putative glycerol uptake facilitator protein, was found to be down-regulated by 60% in C180-2FR at an early growth phase. Two SNPs are located in this gene cluster, one in its promoter region and the other in its protein-coding region. A gene cluster containing genes coding for fluoride antiporters was up-regulated 10-fold in C180-2FR when compared to that in C180-2, independent of growth phase. Expression of the genes containing or in proximity to the SNPs in C180-2 and C180-2FR was then quantified by real-time PCR. Through extensive bioinformatic analysis, we were able to identify 8 single nucleotide polymorphisms (SNPs) in the genome of C180-2FR, which were further confirmed by Sanger sequencing. We performed 50 bp paired-end Illumina shotgun sequencing for both strains. 
The aim of this study is to identify such changes in a fluoride-resistant Streptococcus mutans strain (C180-2FR) using whole-genome shotgun (WGS) sequencing and to examine the potential function of the identified mutations by comparing gene expression between the fluoride-sensitive (C180-2) and C180-2FR strains. However, until now, no studies have reported these genotypic changes. It was hypothesized that these phenotypic differences were due to stable genotypic changes in the fluoride-resistant strains. It is known that fluoride-resistant microorganisms are different from fluoride-sensitive ones in growth, adherence and metabolic activity.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
